News & Events
New release: Mandible vs tongue involvement in cleft palate in mouse models (22 April 2026)
A new dataset is now available on FaceBase from Goodwin, Green, and colleagues at the University of Pittsburgh. Their study examines the respective contributions of mandibular hypoplasia and tongue malposition to cleft palate in Pierre Robin sequence (PRS), using two complementary mouse models. A related manuscript has been accepted for publication in the Journal of Dental Research.
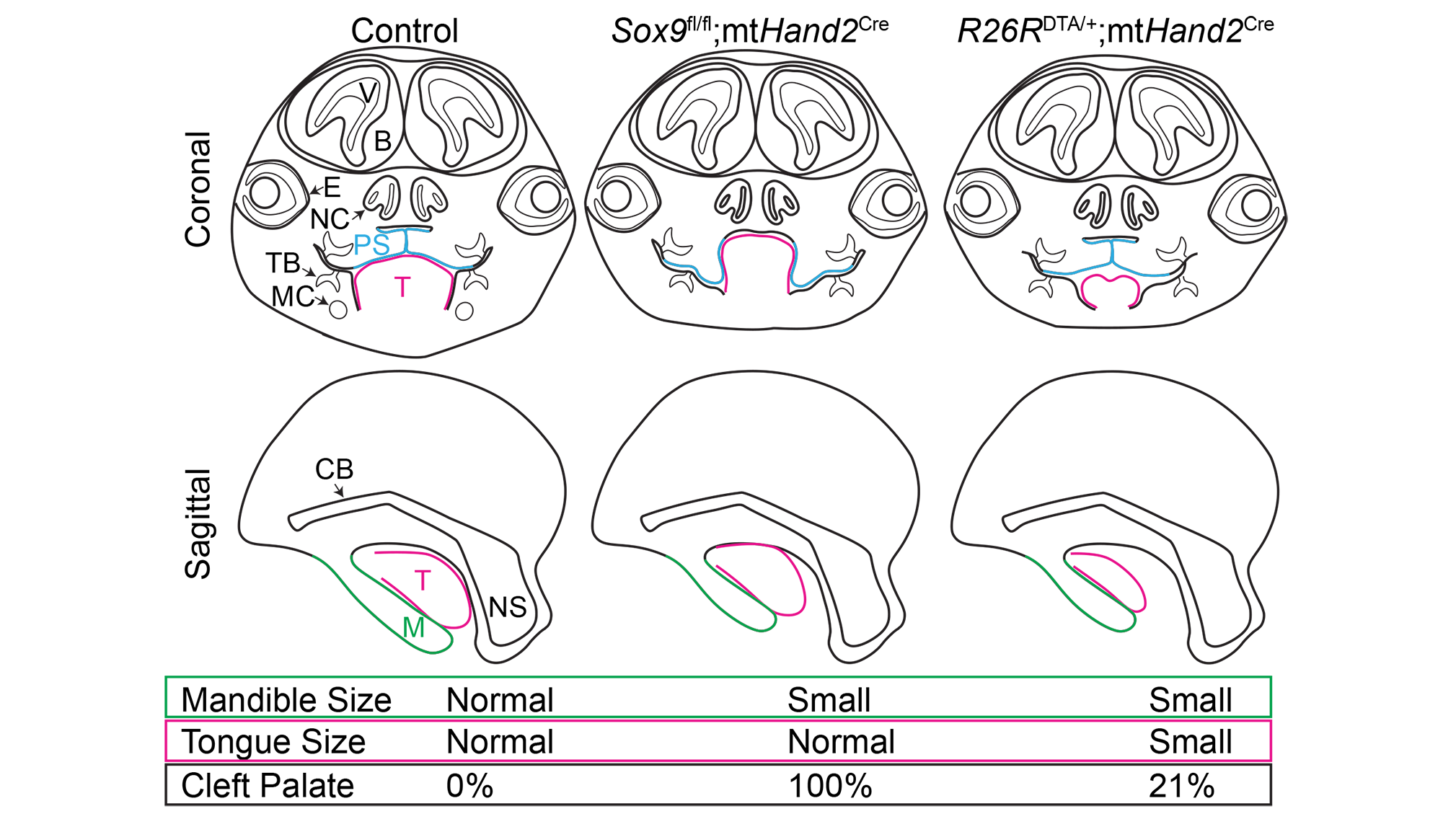 Comparison of palate outcomes in the Sox9fl/fl;mtHand2Cre Pierre Robin sequence model (left) and the DTA/+;mtHand2Cre micrognathia/microglossia model (right). Image courtesy of Alice Fitzgerald Goodwin.
Comparison of palate outcomes in the Sox9fl/fl;mtHand2Cre Pierre Robin sequence model (left) and the DTA/+;mtHand2Cre micrognathia/microglossia model (right). Image courtesy of Alice Fitzgerald Goodwin.
Contributors: Alice Fitzgerald Goodwin, Jeremy Green (University of Pittsburgh)
Description:
To investigate cleft palate in Pierre Robin sequence (PRS), we generated a mouse model with Sox9 deleted specifically in the mandibular mesenchyme (Sox9fl/fl;mtHand2Cre), which resulted in mandibular hypoplasia and retrognathia, palatal shelf elevation delay, and fully penetrant cleft of the secondary palate. To determine the relative contributions of mandible vs tongue malposition to cleft palate in PRS, we generated a micrognathia and microglossia model (DTA/+;mtHand2Cre). The majority of these animals had a normally formed palate, suggesting that tongue obstruction of palatal shelf elevation is the primary contributor to cleft palate in PRS.
Data deposited on FaceBase include microCT scans of Sox9fl/fl;mtHand2Cre and control embryos at E18.5; H&E-stained coronal sections at E12.5, E13.5, E14.5, and E16.5; proliferation and apoptosis assays at E12.5 and E14.5 in the Meckel’s cartilage; immunofluorescence with antibodies against Pax7 and MHC at E14.5; RNAscope with probes against osteogenic markers at E14.5; palatal shelf explant studies at E13.5; and H&E staining and TUNEL apoptosis staining of DTA/+;mtHand2Cre and control embryos.
FaceBase Dataset:
Alice Fitzgerald Goodwin, Jeremy Green. Mandible vs tongue involvement in cleft palate in mouse models. FaceBase Consortium https://doi.org/10.25550/AB-SJQA (2026).
Publication:
Alice Fitzgerald Goodwin, Jeremy Green; Intrinsic tension drives palatal shelf reorientation post tongue retraction. Journal of Dental Research 2026. (accepted April 21, 2026 — DOI to be added upon publication)
New release: Six2 and CTCF ChIP-seq datasets of wildtype mouse embryonic facial tissues (E10.5 - E13.5) (16 April 2026)
New ChIP-seq datasets are now available in FaceBase, profiling Six2 transcription factor binding and CTCF-defined topologically associating domains in wildtype mouse embryonic facial tissues at stages E10.5 through E13.5. Input libraries are included as controls. Together these data support investigation of gene regulatory networks and chromatin architecture during craniofacial morphogenesis.
Contributors: Jingyue Xu, Han Liu and Rulang Jiang (Cincinnati Children’s Hospital, Medical Center)
Description:
We performed a ChIP-seq assay using a Six2 antibody to identify endogenous Six2-binding genomic loci in mouse embryonic facial tissues. We performed a ChIPseq assay using the CTCF antibody to identify topologically associating domains and boundaries. We generated ChIP libraries and sent them for next-generation sequencing. We also generated input libraries as control samples.
FaceBase Dataset:
Jingyue Xu, Han Liu, Rulang Jiang. Six2 and CTCF ChIP-seq datasets of wildtype mouse embryonic facial tissues (E10.5 - E13.5). FaceBase Consortium https://doi.org/10.25550/7Z-41CW (2026).
New release: Phenotypic impacts of embryonic exposure to the CSF1R inhibitor PLX5622 on craniofacial development (09 April 2026)
A new dataset is now available on FaceBase from Ma, Rosin, and colleagues at The University of British Columbia. Their study examines how disrupting CSF1R signaling, which is critical for macrophage and osteoclast development, affects craniofacial morphogenesis during embryonic development. A related manuscript is released on Development.
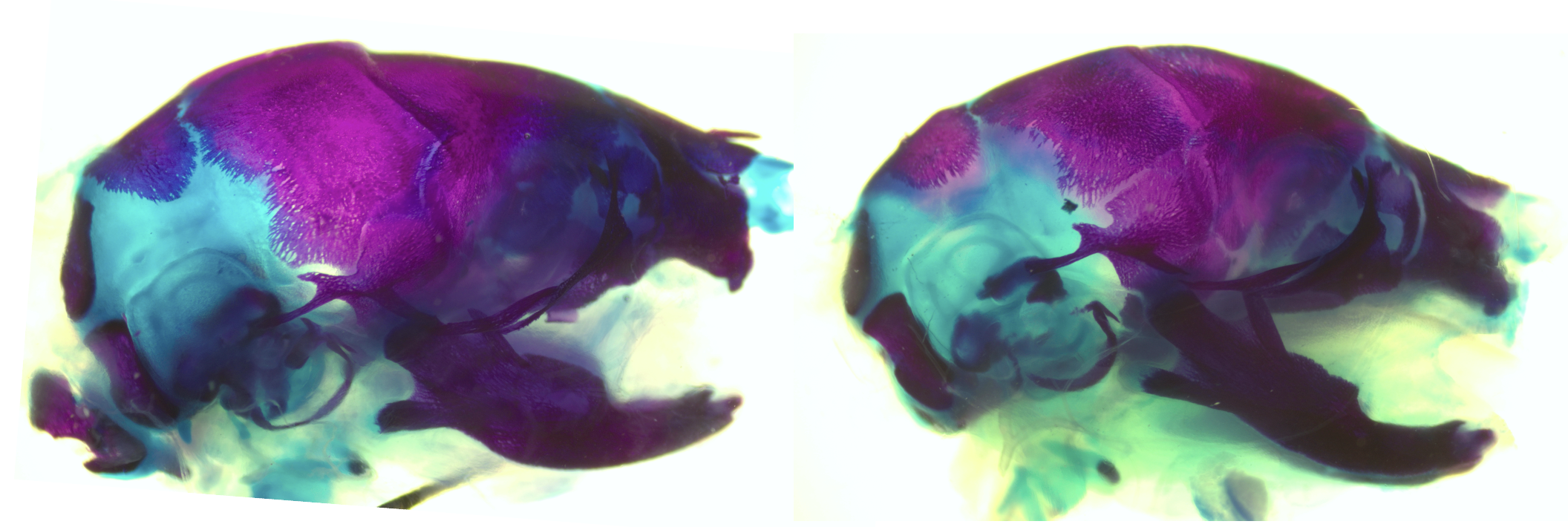 Control (left) and PLX5622-exposed (right) mouse crania stained for bone (red) and cartilage (blue), revealing craniofacial structural disruptions following prenatal CSF1R inhibition. Image courtesy of Felix Ma and Jessica M. Rosin.
Control (left) and PLX5622-exposed (right) mouse crania stained for bone (red) and cartilage (blue), revealing craniofacial structural disruptions following prenatal CSF1R inhibition. Image courtesy of Felix Ma and Jessica M. Rosin.
PI: Jessica M. Rosin (the University of British Columbia)
Description:
Despite a wealth of knowledge on the mechanisms underlying craniofacial morphogenesis during gestation, the roles of fetal macrophages and osteoclasts during this process remain less well characterized. Here, we used the pharmacological inhibitor PLX5622 to disrupt colony stimulating factor-1 receptor (CSF1R) signaling, which is essential for macrophage and osteoclast proliferation, differentiation, and survival. Prenatal PLX5622 exposure resulted in ∼50% depletion of CSF1R+ macrophages, with complete loss of osteoclasts. While there were no notable changes in craniofacial nerve or muscle development, prenatal exposure to PLX5622 resulted in skull doming and cranial suture impairments, in addition to disruptions to development of the premaxilla, mandible, ear ossicles, palate, and cranial base. In response to PLX5622 exposure, cytokine and chemokine signaling was altered and neural crest proliferation was impaired. Our data also highlight sex- and strain-specific differences in PLX5622 phenotypes and together demonstrate that CSF1R+ macrophages and osteoclasts are essential for craniofacial morphogenesis.
Data deposited on FaceBase include images of control and PLX5622-exposed whole-mount E11.5, E12.5, and E13.5 nerve (2H3) and muscle (MF 20) antibody staining and P1 skeletal staining.
FaceBase Dataset:
Felix Ma, Jessica M. Rosin. Phenotypic impacts of embryonic exposure to the CSF1R inhibitor PLX5622 on craniofacial development. FaceBase Consortium https://doi.org/10.25550/A9-12PJ (2026).
Publication:
Felix Ma, Rose Ru Jing Zhou, Matthew Rosin, Iris Zhou, Sabrina Ownsworth, Rouzbeh Ostadsharif Memar, Vincent B. Wong, Jessica M. Rosin; CSF1R+ macrophage and osteoclast depletion impairs neural crest proliferation and craniofacial morphogenesis. Development 2026; dev.205423. doi: https://doi.org/10.1242/dev.205423
Meet FaceBase at GRC and AADOCR/IADR next week (19 March 2026)

FaceBase will be at two major conferences this March. If you’re attending either event in Southern California, we’d love to connect — stop by, say hello, and learn more about our data resources and tools.
Gordon Research Conference
Craniofacial Morphogenesis and Tissue Regeneration
📍 Ventura, CA | March 22–27, 2026
FaceBase co-PI Yang Chai (USC/Ostrow School of Dentistry) will serve as Discussion Leader for the Stem Cells and Organoids I: Stem Cells of the Craniofacial Complex session on Thursday, March 26 (9:00 AM – 12:30 PM).
Alejandro Bugacov and Rob Schuler (ISI/USC) will host an informal demo session on Tuesday, March 24 at 3:00 PM, where attendees can get a hands-on look at the platform and learn how to find and use FaceBase data and tools.
AADOCR/IADR Annual Meeting
American Association for Dental, Oral, and Craniofacial Research
📍 San Diego, CA | March 25–28, 2026
Visit us in the Exhibit Hall at Booth #402 to explore FaceBase resources, see demos, and connect with our team.
Also:
-
Alejandro Bugacov will provide demos at the NIH Trainee Poster Session
📅 Wednesday, March 25 | 9:00 AM – 12:00 PM (PST)
📍 Rooms CC 30A & B -
Jifan Feng (USC/Ostrow School of Dentistry) will present at the NIDCR Data Science – Multi-omics Integration and Precision Medicine Session on Spatial Transcriptomics and Cell Lineage Analysis in Craniofacial Development 📅 Wednesday, March 25 | 1:30 PM – 3:00 PM (PST)
📍 Room 29B
We look forward to seeing you there.
If you’d like to schedule time with our team in advance, feel free to reach out:
help@facebase.org
New release: Spatial Transcriptomics of Mouse CFA-induced TMJ arthritis (04 March 2026)
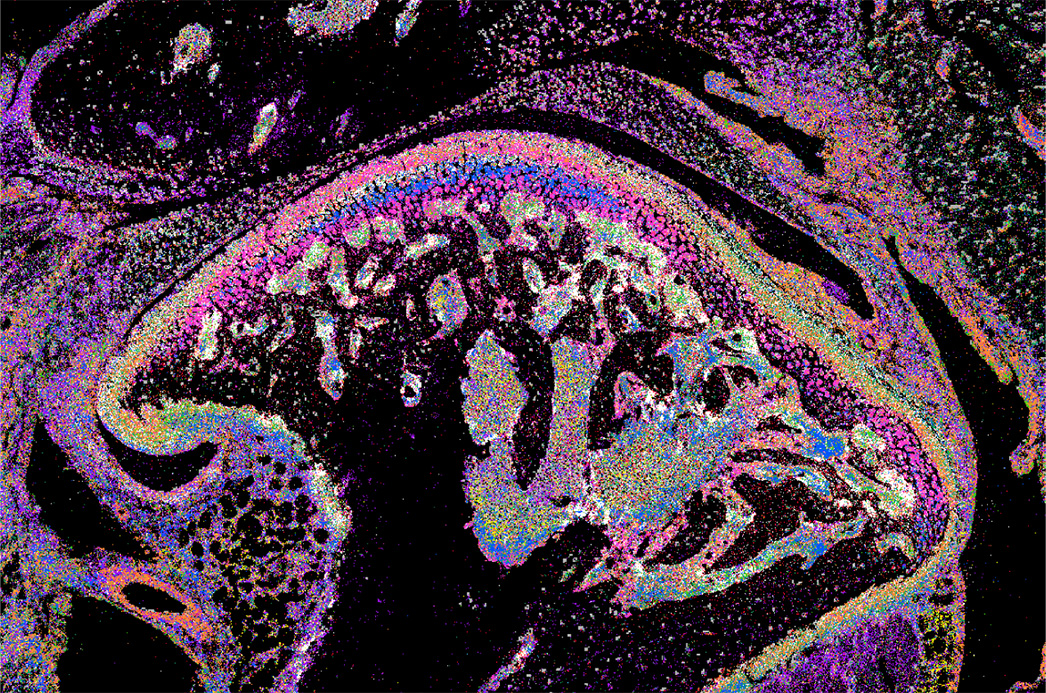 Spatial transcriptomics of the mouse temporomandibular joint (TMJ) using seqFISH, from the dataset Spatial Transcriptomics of Mouse CFA-induced TMJ arthritis
Spatial transcriptomics of the mouse temporomandibular joint (TMJ) using seqFISH, from the dataset Spatial Transcriptomics of Mouse CFA-induced TMJ arthritis
PI: Jian-Fu Chen (University of Southern California)
Description:
Complete Freund’s Adjuvant (CFA)-induced Temporomandibular joint (TMJ) arthritis remodels the cartilage, subchondral bone, and synovial tissue with diverse cell changes. The functional importance of the anatomical organization of TMJ cell types and cellular microenvironment in painful arthritis remains largely unknown. Here, we applied seqFISH (sequential Fluorescence In Situ Hybridization) spatial transcriptomicsto examine the adult mouse TMJ. We uncovered new cell types and comprehensively mapped anatomical locations of diverse cell types with distinct neighborhoods, revealed arthritis-induced cell number and cell status changes, and discovered microenvironment remodeling of fibroblast-immune cells, which are confirmed in patient synovial tissues. Functional and mechanistic studies showed that macrophage-specific knockout of mouse Igf1 promotes its immune activation and upregulates Il33 in adjacent synovial fibroblasts, resulting in inflammatory fibroblast expansion. In turn, fibroblast-specific deletion of Il33 alleviates inflammatory macrophages and inflammation, leading to pain mitigation. Thus, spatial transcriptomics maps diverse cell types in TMJ and reveals a remodeling of synovial fibroblast-immune microenvironment via the Igf1-Il33 axis, which drives arthritis pain with therapeutic potentials.
FaceBase Dataset:
Ziying Lin, Supawadee Jariyasakulroj, Yang Shu, Jingyi Chen, Qing Chang, Pao-Fen Ko, Yuyueyang Qiu, Feixiang Chen, David Ahn, Zhen Zhao, Jian-Fu Chen. Spatial Transcriptomics of Mouse CFA-induced TMJ arthritis. FaceBase Consortium https://doi.org/10.25550/8Q-3KAR (2026).
Publication:
Lin Z, Jariyasakulroj S, Shu Y, Chen J, Chang Q, Ko PF, Qiu Y, Chen F, Ahn D, Zhao Z, Chen JF. Spatial Transcriptomics of TMJ Reveals a Remodeling Fibroblast-Immune Microenvironment Driving Arthritis Pain. Adv Sci (Weinh). 2026 Jan 7:e19816. doi: 10.1002/advs.202519816. Epub ahead of print. PMID: 41498747.
